| Citation: | Long Huang, Lishi Zhang, Dan Li, Rongfei Yan, Weiping Shang, Yunlei Jiang, Shi Li. 2022: Molecular evidence of introgressive hybridization between related species Jankowski's Bunting (Emberiza jankowskii) and Meadow Bunting (Emberiza cioides) (Aves: Passeriformes). Avian Research, 13(1): 100035. DOI: 10.1016/j.avrs.2022.100035 |
Natural hybridization, which often occurs between closely related species exhibiting sympatric or parapatric distributions, is an important source of genetic variation within populations. The closely related Jankowski's Bunting (Emberiza jankowskii) and Meadow Bunting (E. cioides) are similar in morphology and genetics, occupy overlapping niches, and are sympatric in eastern Inner Mongolia. Previous studies have reported trans-species polymorphisms of alleles between the two species, as well as an unexpectedly high genetic diversity of the endangered E. jankowskii. We speculate that introgressive hybridization has occurred between the two species and contributed to the additional unexpected variation to E. jankowskii. We used mitochondrial NADH dehydrogenase subunit 2 (ND2) gene and 15 nuclear microsatellite markers to compare the genetic diversity of E. jankowskii and E. cioides, and inferred the origin of trans-species polymorphisms between the two species by phylogenetic reconstruction and Bayesian cluster analysis. The two species could be clearly distinguished by population cluster analysis. Despite the large number of mutational differences, we still detected sharing of major haplotypes and the presence of hybrids between the two species. Our study confirmed that weak introgressive hybridization has occurred between sympatric E. jankowskii and E. cioides, which may be mediated by female E. cioides individuals, and that interspecific introgression has contributed to the maintenance of high genetic diversity in E. jankowskii. While being wary of the potential negative effects of introgressive hybridization, we suggest that expanding the habitat of E. jankowskii remains the most effective conservation strategy at present.
Obligate avian brood parasites lay their eggs in the nests of other species to impose the burden of parental care; consequently, the hosts rear alien chicks at the expense of their own progeny (Rothstein, 1990; Davies, 2000). Approximately 100 species of birds, accounting for 1% of all bird species, are known to be obligate brood parasites (Payne, 1977; Davies, 2000), and most are generalists with regard to host selection, that is, individuals of the same brood parasitic species may exploit different host species. In general, the degree of host specialization, which can be defined at the species level as the number of host species, phylogenetic relatedness between hosts, or functional diversity (e.g., body size, nest type) of hosts, greatly varies among brood parasitic species or lineages (Davies, 2000; Medina and Langmore, 2016). For instance, the Common Cuckoo (Cuculus canorus) is known to parasitize the nests of over 100 host species (Wyllie, 1981; Moksnes and Røskaft, 1995; Davies, 2000; Rothstein et al., 2002), whereas the Oriental Cuckoo (Cuculus optatus, a close relative to the Common Cuckoo) is known to exploit less than 30 host species (Payne, 2005; Lowther, 2018). In South Korea, for example, the eggs of the Common Cuckoo were found in the nests of six different species (mainly Sinosuthora webbiana, Phoenicurus auroreus, and Emberiza cioides), whereas the Oriental Cuckoo is known to exploit three host species (mainly Phylloscopus coronatus) (Lee, 2014; Lee et al., 2014). This variation has also been observed at the genus level (Jin et al., 2020). For example, the average number of host species for the genus Cuculus is approximately 50, but the genus Hierococcyx exploits only 12 host species (Stokke et al., 2018; Lowther, 2018; Jin et al., 2020). Several factors, such as brood parasite body size, parasite virulence, species age (i.e., time of speciation), and geographic location and range, have been proposed to explain the variations in host specialization among parasitic species (Payne, 1977; Jin et al., 2020; Yang et al., 2020), although we could not completely rule out the possibility of biased study effort among species and/or higher levels. Recently, Medina and Langmore (2016) evaluated a total of 34 parasitic species from four different families (Cuculidae, Icteridae, Indicatoridae, and Viduidae) and suggested that smaller and evolutionary younger species display much narrower host specialization than larger and evolutionary older species. They also showed that the number of hosts was positively associated with the geographic range of the brood parasites. However, the explanatory power of these factors is likely to be inconsistent, depending on the lineage of brood parasites and the selection of specialization index (i.e., the number of host species, phylogenetic relatedness among hosts, or functional diversity).
For an evolutionary young brood parasitic species of host specialists to become an older, generalist species, some pioneering individuals must be ready to attempt parasitic egg-laying in the nests of unfamiliar, new species or be willing to explore a new area in search of hosts. These propensities are likely to be affected by individual attributes, such as individual consistency (i.e., personality) or plasticity (Sinn et al., 2008; Dingemanse et al., 2010; Stamps and Groothuis, 2010), although these two attributes may not be mutually exclusive (Dingemanse et al., 2010). Personality refers to consistent differences (i.e., average difference) in behavior among individuals across a spectrum, which could be expressed as, for example, shy/bold or active/inactive. Meanwhile, individual plasticity represents a change in an individual's responsiveness (e.g., behavior) to external stimuli, such as environmental variation (i.e., variance difference) (Sih et al., 2004; Frost et al., 2007; Ord et al., 2010; Biro and Adriaenssens, 2013; Jolles et al., 2019; Couchoux et al., 2021). It has been well recognized that individual animals differ in the characteristics of both personality and degree of plasticity (Dingemanse et al., 2010; Réale and Dingemanse, 2010). Many experimental studies have shown that such variations in individual attributes, determined via various experimental setups, are often closely associated with other life-history traits and behaviors in the wild, such as individual survival (Hall et al., 2015), dispersal tendency (Dingemanse et al., 2003), prey selection (i.e., accepting a novel prey) (Exnerová et al., 2010), and cuckoo egg rejection by host species (Zhang et al., 2021), providing examples of behavioral syndromes (Sih et al., 2004).
Circumstantial evidence suggests that any variations in individual attributes among parasitic species may contribute to varying degrees of host specialization (Cote et al., 2010; Fogarty et al., 2011). For example, given the positive association between the number of hosts and geographic ranges of the parasite (Payne, 1977; Jin et al., 2020), we may expect that bolder, more exploratory, and risk-taking individuals or species predominantly containing such individuals might be more likely to expand their ranges through the increased dispersal performance of such individuals (Dingemanse et al., 2003; Doligez et al., 2009; Quinn et al., 2011), and thus have a greater chance of including a new host species than parasitic species with opposing characteristics (Medina and Langmore, 2016). Furthermore, we may also expect that such individuals may more readily lay eggs in the nests of novel host species, as similarly seen in prey selection, in which, for example, bolder and more aggressive individuals attack novel insects more frequently and readily (Exnerová et al., 2010). Collectively, from these behavioral syndrome studies, we may expect that the individual attributes of brood parasitic species may influence the chance to encounter and consequently include a new host species. Despite these potential relationships, intra- and inter-specific variations in individual attributes have rarely been considered in the context of host specialization in avian brood parasitism.
In this study, we compared the behavioral performance of two wild-captured Cuculus cuckoos (the Common Cuckoo and the Oriental Cuckoo) in a cage according to the degree of host specialization. These two parasitic cuckoos differ greatly in their number of host species and their breeding ranges (Lee et al., 2014). The Common Cuckoo is widely distributed throughout the Eurasian continent, from the British Isles to the Kamchatka Peninsula in the Russian Far East, during the breeding season, exploiting more than 100 host species (Davies, 2000; Erritzøe et al., 2012). In contrast, the breeding range and number of host species of the Oriental Cuckoo are much smaller than those of the Common Cuckoo, mainly breeding in the Asian region of the Eurasian continent with 26 known host species (Lee, 2014; Lowther, 2018). We quantified the exploratory behavior (e.g., movement) of these species within a birdcage, which is frequently measured to determine individual attributes (e.g., personality), and measured their response (e.g., change in movement) to an experimentally introduced novel object (Boogert et al., 2006; Herborn et al., 2010; Greggor et al., 2015; Costa et al., 2020). We also evaluated their behavioral performance with regard to brood parasites and their number of hosts, testing the prediction that the Common Cuckoo, which has many host species, is more exploratory in a cage than the Oriental Cuckoo. We then suggest future investigative directions to improve our understanding of the role of individual attributes in the variation of host specialization among avian brood parasites.
Fieldwork to catch wild cuckoos was conducted during two breeding seasons (April–June 2018 and 2019) in and around Yangpyeong-gun, Gyeonggi-do, South Korea (37°34′ N, 127°20′ E). The cuckoos were captured using mist nets with the playback of male and female calls of the respective species and 3D-printed decoys that mimic each species. Over the two years, we captured 49 Common Cuckoos (43 males and 6 females) and 19 Oriental Cuckoos (12 males and 7 females), with which we conducted the cage experiments (see below for details). After the experiments, all cuckoos were measured (wing, tarsus, bill, and mass) and metal-ringed for individual identification. They were then safely released at the capture site.
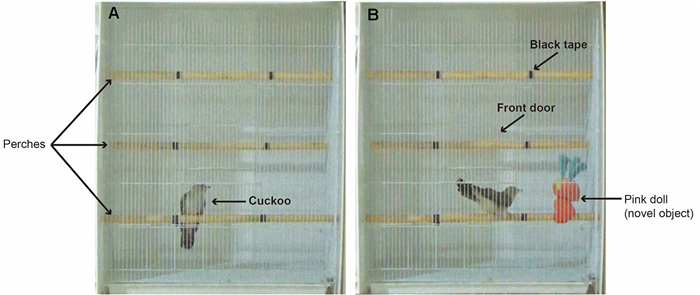
Upon capture, we placed each cuckoo in a bird bag and transported it by car (approximately 30 min) to an indoor garage located in the middle of the study area where the cage experiments were conducted. To quantify behavioral performance, we followed the experimental design and procedures developed by Kluen et al. (2012) and reliably applied to other birds (e.g. Herborn et al., 2010; Friard and Gamba, 2016; Zhang et al., 2021), with some modifications in terms of cage size and experiment time. The size of the cage was 900 mm (height) × 760 mm (width) × 510 mm (depth), and all sides except the front were covered with white foam boards (Fig. 1). Three perches were mounted at the bottom, middle, and uppermost levels of the cage, each of which was divided into three equal-sized zones using black tape (Fig. 1). Accordingly, we divided the cage vertically into four zones (i.e., the uppermost, middle, and low perches, and the bottom of the cage) and horizontally into three sections (left, middle, right), thereby generating 12 hypothetical zones in the cage. We also included the front wires as a zone, resulting in 13 zones used to locate the bird in the cage. The lighting condition of the garage was consistent throughout the experiments, and the researcher left the garage and kept the door closed during the experiment to minimize disturbance.
The experimental procedures for each bird followed the same protocol. Each bird was released into the cage and allowed to habituate for 10 min without disturbance. After the acclimation period, the novel object and control experiments were conducted successively in a random order. For the novel object experiment, we randomly selected a location among the four zones defined by perches (i.e., the leftmost and rightmost zones of the uppermost and lowest perches, Fig. 1) to place a novel object (a pink doll with no eyes), and then recorded the behavior of the bird for 3 min. For the control experiment, we recorded bird behavior for 3 min in the absence of the novel object prior to or after the novel object experiment, at random. When the control experiment was performed first, following the protocol of Kluen et al. (2012), the researcher placed one hand in the cage through the front door and moved it once or twice as if placing the doll on a perch before recording. When performed second, we recorded the behavior after removing the doll from the cage. All experimental procedures were recorded using a SONY HDR-PJ340 Handycam camcorder (SONY KR) installed approximately 2 m in front of the cage. Due to ethical reasons and logistical constraints for breeding birds, we conducted the experiment once for each bird, although this may hinder us from reliably measuring the repeatability of their behavior.
Using the Behavioral Observation Research Interactive Software (BORIS, version 6.3.9) (2016), we extracted the following behavioral data from each video to evaluate their levels of exploratory behavior: movement (number of movements between zones defined in the cage), head-turning (number of head turns), wing-flapping (number of wing flaps), and stepping (number of steps, including hops, on the bottom of the cage and perches). We also determined the location of the birds in the cage based on the defined zones and measured the approach time (in seconds) toward the novel object within the distance of individual body length as an indicator that may express the boldness and exploration of individuals (Wilson et al., 1993; Frost et al., 2007; Stamps and Groothuis, 2010). To minimize potential interference caused by the progress of experiments, we excluded the first 30 s after the start of each test (180 s) from the analysis and extracted behavioral data from the remaining 150 s.
The relationships between the four behavioral types (movement, head-turning, wing-flapping, and stepping) of two Cuculus cuckoos were measured using the Spearman rank correlation test. Behavioral differences between the control and experimental groups of a species, the effect of their orders, and the interspecific differences were also evaluated using the Wilcoxon signed-rank test. For the four behavioral types in the two species, we conducted principal component analysis (PCA) to evaluate their exploratory behavior with a reduced, uncorrelated dataset. Principal components (PCs) were chosen by the cumulative percentage of total variation, for which we chose the smallest number of components that accounted for more than 80% of total variance (Jolliffe, 2002). Principal components were compared between the two species and between the sexes, using the two-sample t-test or Wilcoxon rank sum test, depending on the nature of data. All statistical analyses were conducted using R version 4.0.3 (R Core team, 2020).
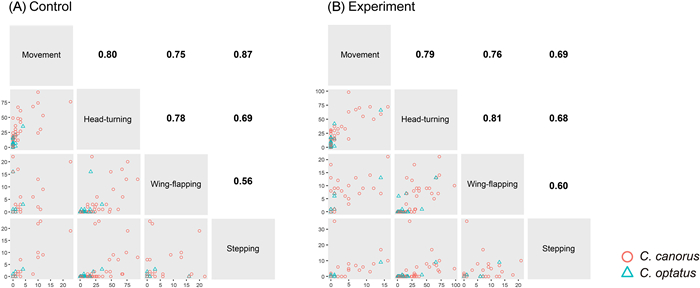
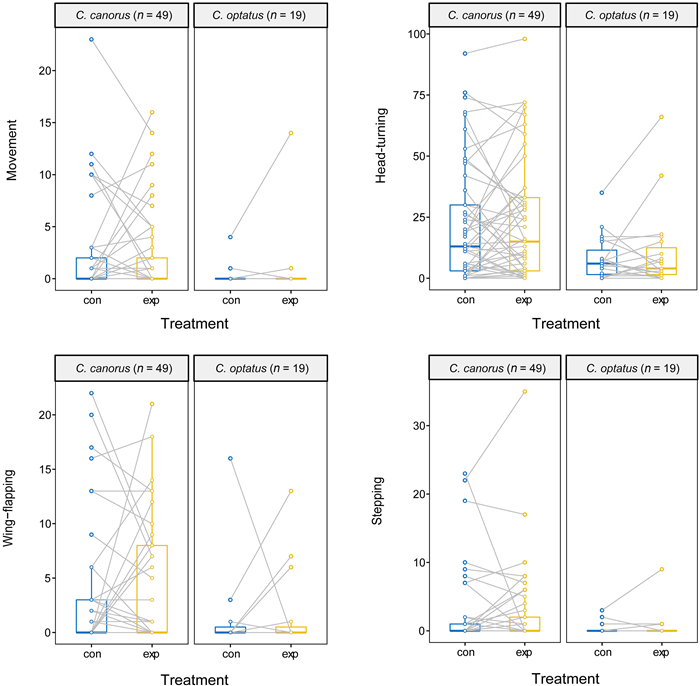
In general, the Common Cuckoo tended to exhibit higher activity on average across the four behavioral types than the Oriental Cuckoo in both treatments, with various significance levels (Table 1), and the similar results were emerged when we analyzed with males only (Appendix Table S1). In both species, most of the four behaviors were highly correlated with one another (ranges of rs: Common Cuckoo 0.56–0.87, Oriental Cuckoo 0.14–0.56, Fig. 2a; Common Cuckoo 0.60–0.81, Oriental Cuckoo 0.46–0.90, Fig. 2b). We also found that the average and variance of all types of behavioral activity were not significantly different between the control and experimental groups of the two species, with treatment order having no effect (all p > 0.05; Fig. 3). Notably, these similarities appeared to be caused by random changes in individual behavior between treatments, rather than personality, indicating an inconsistent effect of the novel object on an individual's behavior. Regarding approach behavior, only five Common Cuckoos and no Oriental Cuckoos approached the novel object within the distance of their body size, with time ranging from 17.2 to 139.2 s. However, it is unclear whether these cuckoos intentionally approached the object, or whether the approach was made as a result of random movement in the cage.
| Control | Experiment | ||||||||
| C.canorus | C.optatus | W | p | C.canorus | C.optatus | W | p | ||
| Movement | 2.24 | 0.37 | 586 | 0.03 | 2.20 | 0.89 | 561 | 0.06 | |
| Head-turning | 22.61 | 8.21 | 621 | 0.02 | 23.27 | 10.79 | 615 | 0.02 | |
| Wing-flapping | 2.94 | 1.16 | 558.5 | 0.07 | 3.90 | 1.47 | 573 | 0.05 | |
| Stepping | 2.35 | 0.32 | 542.5 | 0.09 | 2.41 | 0.58 | 552 | 0.07 | |
Principal component analysis (PCA) was applied to examine the overall level of exploratory behavior of the two cuckoo species in the cage according to the experimental conditions (i.e., control and experiment). In the control group, the first two PCs accounted for more than 80% of the total variance in the data (Table 2). PC1, which explained 70% of the total variation, included movement and head-turning behavior, the level of which increased with increasing PC values. Meanwhile, PC2 accounted for 20% of the total variation and best explained the variability of wing-flapping and stepping, and the two components exhibited opposite relationships with increasing PC values (i.e., higher PC values represent increasing wing-flapping and decreasing stepping, and vice versa). In the experimental group, the first two PCs explained 88% of the total variance in the data (Table 2). As in the control group, PC1 accounted for 70% of the total data, including movement, head-turning, and wing-flapping, the level of which increased with increasing PC values. In addition, stepping best explained the data in PC2, the level of which increases with increasing PC values.
| Variables | Control | Experiment | |||
| PC1 (70%) | PC2 (20%) | PC1 (70%) | PC2 (18%) | ||
| Movement | 0.56 | −0.28 | 0.55 | −0.03 | |
| Head-turning | 0.52 | 0.28 | 0.55 | −0.17 | |
| Wing-flapping | 0.43 | 0.70 | 0.50 | −0.45 | |
| Stepping | 0.49 | −0.60 | 0.38 | 0.88 | |
| Total explanation power 90/88%. | |||||
We observed that the Common Cuckoos exhibited significantly larger PC1 values than that of the Oriental Cuckoos in the control group (t = 2.18, p < 0.05). The proportion of individuals that had exceptionally large PC 1 value (e.g. 2 or more, so much more exploratory) was much larger in the Common Cuckoo: about 16% (8/49) of Common Cuckoos having PC1 values of 2 or more but none (0/19) in the Oriental Cuckoo. Similarly, PC1 values were marginally significantly different in the experimental group (t = 1.93, p = 0.06), in which about 20% of the Common Cuckoos had a PC 1 value of 2 or more but only one (5%) individual in the Oriental Cuckoo. These results indicate that the Common Cuckoo generally tended to exhibit more exploratory behavior in the cage than the Oriental Cuckoo, irrespective of experimental treatment (Fig. 4). However, there were no clear differences in PC2 between the two species in either treatment group. We also found a sexual difference in PC1 of the experimental group in the Common Cuckoo (w = 74, p < 0.05), in which males had higher PC1 value, but not others (all p > 0.05).
This study was conducted, for the first time to our knowledge, to compare the exploratory behavior of two Cuculus species of cuckoos that have variable degrees of host specialization. Behaviors exhibited by the cuckoos in the cage were classified into four types: movement between defined zones in the cage, head-turning, wing-flapping, and stepping. In the Common Cuckoo, the occurrences of these behaviors were highly correlated with one another, that is, individuals with more movement also exhibited more stepping as well as wing-flapping and head-turning. Our results show that the Common Cuckoo tended to exhibit higher performance in all types of behavior than the Oriental Cuckoo, and also comprised more individuals that showed highly exploratory behavior, which were retained, regardless of whether the novel object was present in the cage. These results fit with our prediction that the Common Cuckoo, which has more host species, should be more exploratory or include more such individuals than the Oriental Cuckoo. Thus, it is consistent with the hypothesis that the interspecific variation of behavioral characteristics, such as personality and plasticity, may contribute to the varying degree of host specialization among avian brood parasites. As mentioned above, more exploratory, active, and bolder species or species including more of such individuals (here, the Common Cuckoo) may be or become host generalists more readily than those with opposing characteristics as they can expand their ranges with increased dispersal tendency or make more attempts to include a new host species with a neophilic nature. How individual plasticity contributes to this was not tested explicitly in this study and need to be addressed in the future study.
The observed differences in individual attributes between species may also be inferred indirectly from their responses to playback while attempting to catch them and the resultant number of birds captured. The cuckoos attracted to the playback of female calls usually flew around the catching spot where mist nets were set up with the decoy and the loudspeaker, and then attempted to approach the decoy. Here, there were large variations in the approaching time after they were attracted between and within species. In the wild, female cuckoos usually generate calls infrequently; therefore, continuous and repeated playback sounds may provide male cuckoos with unusual and novel environments. As such, any differences in the response to playback may be partly derived from their personality differences. In this study, the Common Cuckoo exhibited the best response to playback, vigorously approaching the decoy to attempt to copulate (Lee et al., 2019). In contrast, the response of Oriental Cuckoos to playback tended to be relatively much more cautious and prudent, which resulted in low catching success and a smaller sample size.
Generally, we found that the two cuckoo species were more inactive in the cage than other birds tested in similar experiments. For example, according to Dingemanse et al. (2002), the Great Tit (Parus major) achieved an average of 20 exploratory scores (i.e., total number of flights and hops) in 2 min. In another case, the Common Starling (Sturnus vulgaris) achieved average exploratory scores of 54.53 for stepping and 40.95 for flight during an 8-min experiment (Dingemanse et al., 2002). In comparison, during this study, the Common Cuckoo achieved average scores of 2.34 for wing-flapping and 2.94 for stepping during the control treatment, and many individuals did not move between zones, only exhibiting head-turning or no movement at all. Furthermore, we observed no tendency toward either neophilia or neophobia through the novel object experiment; they appeared indifferent to the novel object introduced to the cage. The overall low activity of cuckoos in the cage may be associated with their cryptic and secretive nature, which is necessary to facilitate the success of brood parasitism, as is well recognized in the literature (Wyllie, 1981; Davies, 2000; Minderman et al., 2009). However, constraints derived from experimental design, such as insufficient acclimation time or spatial limitations, cannot be ruled out. In a similar context, we may expect sexual differences in individual attributes as females perform egg-laying in avian brood parasites. In this study, however, we only found that female Common Cuckoos exhibited significantly lower exploratory behavior than males in the experimental group, but given small sample size for female cuckoos, further studies are necessary to verify sexual differences in individual attributes.
The results of this study indicate that individual attributes could vary, albeit on a small scale, within as well as among parasitic species, but there were several limitations in this study that need to be addressed in subsequent studies. First, a proper experimental design to determine the behavioral characteristics of medium-sized non-passerine birds should be developed. In this study, we adopted the cage experiments first developed by Kluen et al. (2012) with some modifications because of their feasibility for application in the field with minimum adverse effects on breeding birds, based on the time required for experiments. However, the overall low activity of cuckoos may indicate that further modification of the experimental methodology or designing a new experimental methodology is necessary to more accurately quantify the behavioral characteristics of these avian species. Second, individual attributes may play a role in causing variations in the degree of host specialization, as we suggested here, as well as in the overall process of brood parasitism (such as host nest selection, parasitism frequency, and response to host mobbing). Therefore, studies directly testing the causal effect of individual attributes on these features and its heritability need to be conducted across a range of brood parasites (such as cowbirds) to verify our hypothesis. These future studies will broaden our understanding of the consequences of individual attributes on the coevolution of avian brood parasites and their hosts.
JWL conceived the study; SJJ, GWB, HNK and JWL performed field experiments; SJJ and JWL analyzed data; SJJ, JWL and JCY wrote the manuscript. All authors read and approved the final submission.
Cuckoos were captured with permits by Yangpyeong-gun County, and the fieldwork and experimental procedures conformed to the national law of Korea and the guidelines of the Kyung Hee University Animal Ethics Committee.
The authors declare the following financial interests/personal relationships which may be considered as potential competing interests: Jin-Won Lee reports financial support was provided by National Research Foundation of Korea.
We thank Hye-Kyoung Moon and Myun-Sik Kim for their valuable help during the fieldwork and cage experiment. This research was supported by the National Research Foundation of Korea (NRF-2017R1D1A1B03030329, NRF-2019K2A9A2A06022677).
Supplementary data to this article can be found online at https://doi.org/10.1016/j.avrs.2022.100028.
| Control | Experiment | ||||||||
| C.canorus | C.optatus | W | p | C.canorus | C.optatus | W | p | ||
| Movement | 2.24 | 0.37 | 586 | 0.03 | 2.20 | 0.89 | 561 | 0.06 | |
| Head-turning | 22.61 | 8.21 | 621 | 0.02 | 23.27 | 10.79 | 615 | 0.02 | |
| Wing-flapping | 2.94 | 1.16 | 558.5 | 0.07 | 3.90 | 1.47 | 573 | 0.05 | |
| Stepping | 2.35 | 0.32 | 542.5 | 0.09 | 2.41 | 0.58 | 552 | 0.07 | |
| Variables | Control | Experiment | |||
| PC1 (70%) | PC2 (20%) | PC1 (70%) | PC2 (18%) | ||
| Movement | 0.56 | −0.28 | 0.55 | −0.03 | |
| Head-turning | 0.52 | 0.28 | 0.55 | −0.17 | |
| Wing-flapping | 0.43 | 0.70 | 0.50 | −0.45 | |
| Stepping | 0.49 | −0.60 | 0.38 | 0.88 | |
| Total explanation power 90/88%. | |||||