| Citation: | Xuan Li, Yuxin Zhang, Hongfeng Zhao. 2024: Endochondral ossification of hindlimbs in embryonic development of Japanese Quail (Coturnix japonica). Avian Research, 15(1): 100152. DOI: 10.1016/j.avrs.2023.100152 |
The endochondral ossification of hindlimb is essential to a bird's ability to stand, walk and fly. Most hindlimb is ossified in the embryos before hatching in precocial birds. However, the molecular mechanisms of hindlimb ossification in birds is still unclear. Therefore, we tried to examine the process of hindlimb ossification and its molecular regulation by using an animal model—Japanese Quail (Coturnix japonica). We selected four critical stages (Embryo Day: E6, E8, E12 and E16) of skeletal development of embryonic quails for hindlimb skeleton staining to show the process of endochondral ossification and to examine the molecular regulation of endochondral osteogenesis by RNA-Seq analysis. The results showed that ossification became increased with embryonic development and most hindlimb was ossified before hatching. RNA-Seq analysis revealed that various signaling pathways were involved with endochondral ossification with thyroid hormone signaling and WNT signaling pathway particularly enriched. Moreover, the expression levels of 42 genes were continuously upregulated and 14 genes were continuously downregulated from E6 to E16. The present study might provide new insights into complex molecular mechanisms in regulation of endochondral ossification.
The formation of skeletal system is one of the most important characteristics in vertebrates. Skeletal phylogeny originates from the mesoderm and is composed of cartilage and bone (Yang, 2009). Usually, bone development and growth in birds are accomplished through intramembrane and endochondral ossification. The intramembrane ossification means that mesenchymal cells are directly transformed into bone tissue while endochondral ossification indicates that mesenchymal cells can be differentiated into cartilage templates and then into bones (Ahmed and Soliman, 2013). The bone of hindlimb is mainly shaped by endochondral osteogenesis (Pourlis and Antonopoulos, 2014). For example, bird hindlimb is formed from condensed undifferentiated mesenchymal cells, which then participate in osteogenesis through the proliferation, hypertrophy and maturation of chondrocytes (Ahmed and Soliman, 2013). The process of bird hindlimb ossification is crucial for a bird's ability to stand, walk, and fly.
Japanese Quail (Coturnix japonica) is precocial with well-developed legs and feet after hatching. Its hindlimb endochondral ossification occurs early in embryonic development, helping to facilitate the rapid development of their legs and feet (Maxwell, 2008). This process is tightly regulated by a variety of signaling pathways that control chondrocyte proliferation, differentiation, and hypertrophy, as well as osteoblast differentiation and bone formation. Key signaling pathways involved in endochondral ossification include fibroblast growth factor (FGF) pathway, the Indian hedgehog (IHH) pathway, which controls chondrocyte proliferation and differentiation; the bone morphogenetic protein (BMP) pathway, which promotes osteoblast differentiation and bone formation; and the Wingless/Integrated (WNT) pathway, which regulates both chondrocyte and osteoblast differentiation (Wuelling and Vortkamp, 2010; Nishimura et al., 2012; Bo et al., 2018; Gao et al., 2018).
In recent years, RNA sequencing (RNA-Seq) has become a powerful tool used to study gene expression and transcriptomics in various organisms (Wang et al., 2009). This technique involves the sequencing of RNA molecules present in a sample, which provides valuable information about the identity and abundance of expressed genes (Ozsolak and Milos, 2011). Moreover, Japanese Quail develops rapidly and have well-defined stages of embryonic development, making them a useful tool for studying the cellular and molecular mechanisms that govern avian development (Ainsworth et al., 2010). In addition, the quail genome has been sequenced, facilitating genetic studies aimed at understanding the molecular basis of developmental processes (Morris et al., 2020). As a result, Japanese Quail has been used to study a wide range of developmental phenomena, including organogenesis and limb development (Ainsworth et al., 2010; Pourlis and Antonopoulos, 2014).
Therefore, this study focused on investigating the molecular mechanisms that regulated endochondral ossification during the embryonic development of Japanese Quail by RNA-Seq analysis. The present study might provide new insights into complex molecular mechanisms in regulation of endochondral ossification. Understanding the intricacies of bird embryo hindlimb ossification is important for studying avian development, and understanding this process is important for studying avian biology and behavior, as well as for determining the health and development of individual birds.
The fertilized eggs of yellow-feathered Japanese Quail were purchased from local quail farm. The quail eggs within 6 h after laying were collected and then stored at 13 ℃. Then 100 eggs were randomly selected with an average weight of 11.83 ± 0.75 g and then placed in an automatic incubator with an initial temperature of 38 ℃ and humidity of 60%, and cultured until the 15th day. From the 15th day on, the humidity increases to 70% and the temperature drops to 37.7 ℃. During the whole process of incubation, the eggs were slowly rotated up to a total of 90° every 90 min for normal development (Pourlis et al., 1998; Huss et al., 2008; Baer et al., 2015). Eggs were inspected for embryo viability using a standard egg candler.
On the basis of previous study and skeletal development of quail embryos, embryos were collected on Embryo Day 6 (E6), Day 8 (E8), Day 12 (E12) and Day 16 (E16) of incubation (embryonic development) for skeletal staining and RNA extraction (Pourlis et al., 1998; Nakane and Tsudzuki, 1999). On Day 6 of incubation (E6), ossification center appears in the hindlimb bone and hindlimb is visibly cartilaginous. On Day 8 of incubation (E8), the columnar ossification continues along the diaphysis of all long bones. On Day 12 of incubation (E12), the embryos increase with more ossification in the bone. On Day 16 of incubation (E16), embryonic development completes and is about to hatch. All of animal treatments and experiments were performed in accordance with Animal Care Guidelines of Shaanxi Normal University and Ministry of Science and Technology of the People's Republic of China.
Alizarin red-alcian blue double staining method was used to observe the visual changes of quail embryo bone during its development (Bo et al., 2018). Starting from the 6th day of embryo development, three fertilized eggs were taken out every day, and a hole of 6 mm2 was drilled on the blunt end surface of the eggs with a dentist's drill. After that, the embryos were carefully cut out and immersed in 95% alcohol for fixation for five days. After fixation for 3 h, the organs and skin tissues of the embryos were carefully removed with tweezers. The fixed specimen was bleached with 30% hydrogen peroxide for half an hour and then defatted with acetone, 0.5% KOH was pre-transparent, then stained with a mixture of alizarin red and alcian blue, and finally separated and transparent in a mixture of ethanol and glycerol of different concentrations. Cartilage is stained blue and hard bone is stained red (Gao et al., 2018). After completion, the film was observed under a stereomicroscope equipped with a digital camera. Meanwhile, we used Vernier calipers to measure the length of hindlimb and femur of quail embryo under a dissecting microscope before staining, and then ossification ratio was calculated after staining. The ossification ratio is the percentage of the red region length/length of the femur (Nakane and Tsudzuki, 1999).
On the 6th, 8th, 12th and 16th day (incubation day) of embryonic development, three eggs were taken out respectively, and the hindlimb bones of the embryos were quickly dissected and stored at −80 ℃. Then the bones were ground by tissue grinder homogenizer and total RNA was extracted by using TriZol reagent. The quality and integrity of RNA were evaluated. The cDNA library was constructed and then sequenced through Illumina Novaseq 6000 platform with 150 bp paired sequencing (Ling En, Shanghai, China). Q30 values and guanine-cytosine (GC) content were calculated to evaluate read quality (Wang et al., 2022).
Once the sequencing was complete, the raw data were pruned to obtain a clean sequence, which was then compared with the reference quail genome (GCF_001577835.2) (Morris et al., 2020). The results were used to calculate the fragments per kilobase of exon per million mapped reads (FPKM) value for each transcript in the sample, which indicated the level of gene expression.
To obtain differentially expressed genes (DEGs) during hindlimb development in quail embryos, we compared pair-to-pair samples from four developmental stages. R statistical package edge R (Empirical analysis of Digital Gene Expression in R) was used for differential expression analysis. The differentially expressed genes (DEGs) were identified according to the criteria of |log2 (fold change)| ≥ 1 and false discovery rate (FDR) ≤ 0.05. Go function enrichment and KEGG pathway analysis were performed by Goatools (https://github.com/tanghaibao/Goatools) and KOBAS (http://kobas.cbi.pku.edu.cn/home.do) to understand the function of differentially expressed genes. GO terms and DEGs in the metabolic pathway were significantly enriched when Bonferroni correction P-value < 0.05 (Wang et al., 2022; Zhang et al., 2023).
To verify candidate genes derived from RNA sequencing results, we selected 33 genes involved in hindlimb bone development of quail embryos for qRT-PCR detection. The extracted RNA was reverse-transcribed into cDNA by reverse transcriptase, and β-actin gene was selected as the reference gene to normalize the expression level of target genes. The primer was designed by using Primer 6 and primer sequences were listed in Appendix Table S1. The process of RT-PCR was consistent with previous studies (Zhang et al., 2023). Three biological replicates were performed for each gene sample, and the results were represented by a Bio-Rad CFX Manager software curve. The curve should be a single peak curve with high primer specificity, and the efficiency of all primers should be above 95% after analysis. Finally, we choose 2−ΔΔCT to express the relative expression of genes (Livak and Schmittgen, 2001).
All statistical analysis was conducted in R Package (version 4.2.1). All data were assessed for normality and homogeneity of variance before conducting one-way analysis of variance (ANOVA) with multiple comparison. The results are shown as mean ± standard error (SE) with p < 0.05 considered as significant difference.
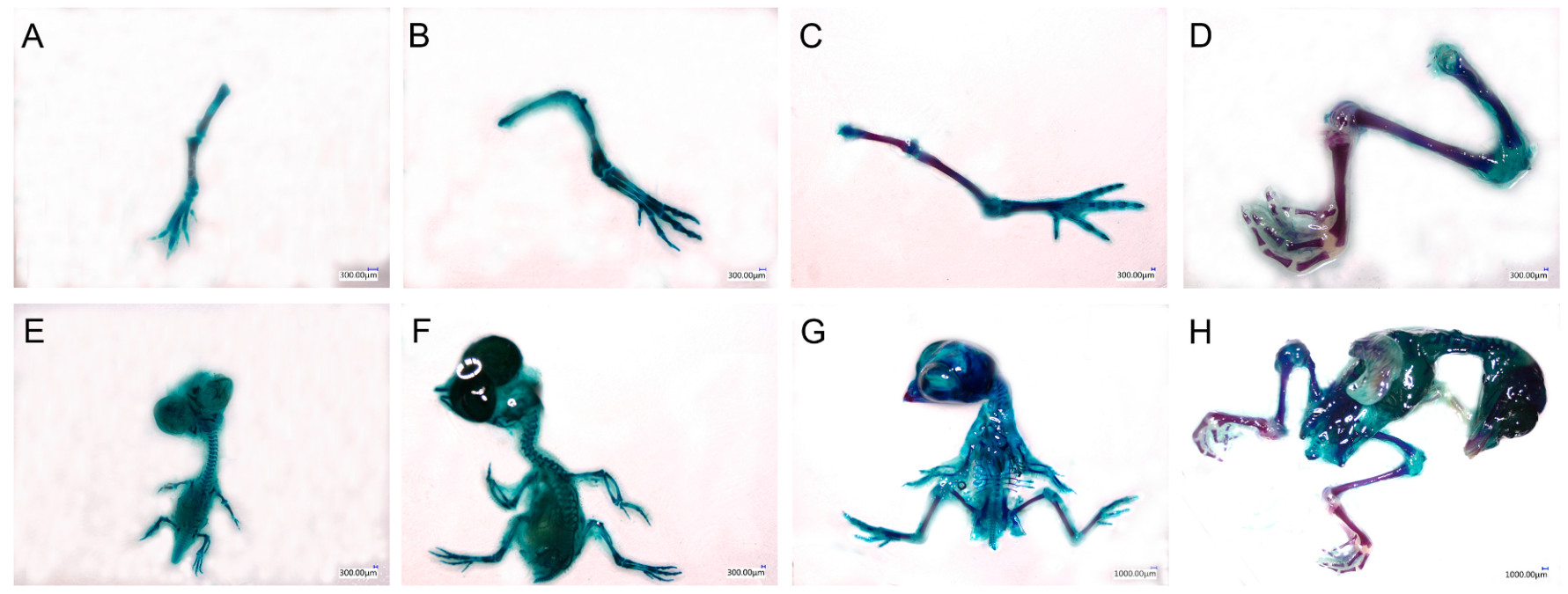
The whole skeleton and hindlimb bone of embryos at E6, E8, E12 and E16 are shown in Fig. 1, and Table 1 demonstrates the length of hindlimb and femur and the ossification of femur in embryos from E6 to E16. At E6, most of skeleton including hindlimb bone was cartilaginous with the ossification portion of femur nearly 20%. At E8, more ossification of whole skeleton was observed. The hindlimb was significantly elongated and the ossification portion was significantly increased in comparison with that in E6. The ossification of whole skeleton and hindlimb including femur was further deepened at E12 and more complete at E16. Significant increase in length and ossification ratio of femur was found in embryos at E12 and E16 (Table 1).
| Developmental stage | Femur length (mm, mean ± SE) | Hindlimb length (mm, mean ± SE) | Ossified proportion of femur (%) |
| E6 | 1.64 ± 0.28 | 9.31 ± 0.06 | 20.6 ± 2.2 |
| E7 | 3.15 ± 0.25 | 11.57 ± 0.13 | 28.6 ± 2.6 |
| E8 | 3.82 ± 0.14 | 12.81 ± 0.13 | 32.8 ± 2.4 |
| E9 | 5.27 ± 0.27 | 20.14 ± 0.34 | 52.6 ± 3.6 |
| E10 | 5.99 ± 0.12 | 23.44 ± 0.18 | 61.4 ± 1.6 |
| E11 | 6.39 ± 0.38 | 29.36 ± 0.14 | 64.2 ± 2.2 |
| E12 | 6.98 ± 0.04 | 33.64 ± 0.27 | 70.8 ± 2.4 |
| E13 | 9.33 ± 0.31 | 40.04 ± 0.06 | 75.2 ± 3.2 |
| E14 | 10.57 ± 0.06 | 44.17 ± 0.14 | 79.4 ± 2.2 |
| E15 | 11.78 ± 0.12 | 51.16 ± 0.39 | 81.6 ± 3.4 |
| E16 | 13.27 ± 0.05 | 54.81 ± 0.38 | 86.2 ± 1.8 |
We sequenced cDNA libraries from four key stages of embryonic bone development of quail. Each sample obtained an average of 57, 446, 137 raw reads, and 51, 979, 691 clean reads after data filtering, with a match rate of more than 86%. We used clean reads for further analysis. Meanwhile, the quality of cleaning data was evaluated to ensure that GC content was above 48% and Q30 content was above 95% (Table S2).
Six groups of DEGs were obtained by pairwise comparison of four stages during embryonic skeletal development (E6 vs E8, E6 vs E12, E6 vs E16, E8 vs E12, E8 vs E16, E12 vs E16). There were 722, 2968, 4930, 2234, 4169 and 2475 DEGs during four stages respectively (E6 vs E8, E6 vs E12, E6 vs E16, E8 vs E12, E8 vs E16, E12 vs E16). Venn diagram showed that a total of 2829 and 2602 DEGs were significantly up-regulated and down-regulated in E6 vs E8, E8 vs E12 and E12 vs E16 (Fig. 2). Among 5431 genes during these four stages, the expression levels of 42 genes were continuously up-regulated and 14 genes were continuously down-regulated.
Then we clustered the expression patterns of differentially expressed genes in hindlimb development of quails into 10 clusters, which represented the locus of increased or decreased expression of these genes during development. Clusters B, J, I and H were specifically expressed in E6, E8, E12 and E16, respectively (Fig. 3). There are 103, 91, 818 and 718 genes that are highly expressed in E6, E8, E12, and E16.
Through search of GO database, gene ontology (GO) enrichment analysis of DEGs in E6 vs E16 group was carried out to further understand the function of these genes, which were classified as biological processes (BP), cell component (CC), and molecular functions (MF) (Fig. 4).
Besides, DEGs in the E6 and E16 groups were localized to 291 KEGG pathways. These KEGG pathways can be divided into cellular processes, genetic information processing, biological systems and environmental processes information processing and metabolism. Among them, the metabolic pathways involved most genes. Figure 5 shows top 30 significantly enriched metabolic pathways. In addition, WNT signaling pathway and thyroid hormone signaling pathway were also shown in Fig. 5.
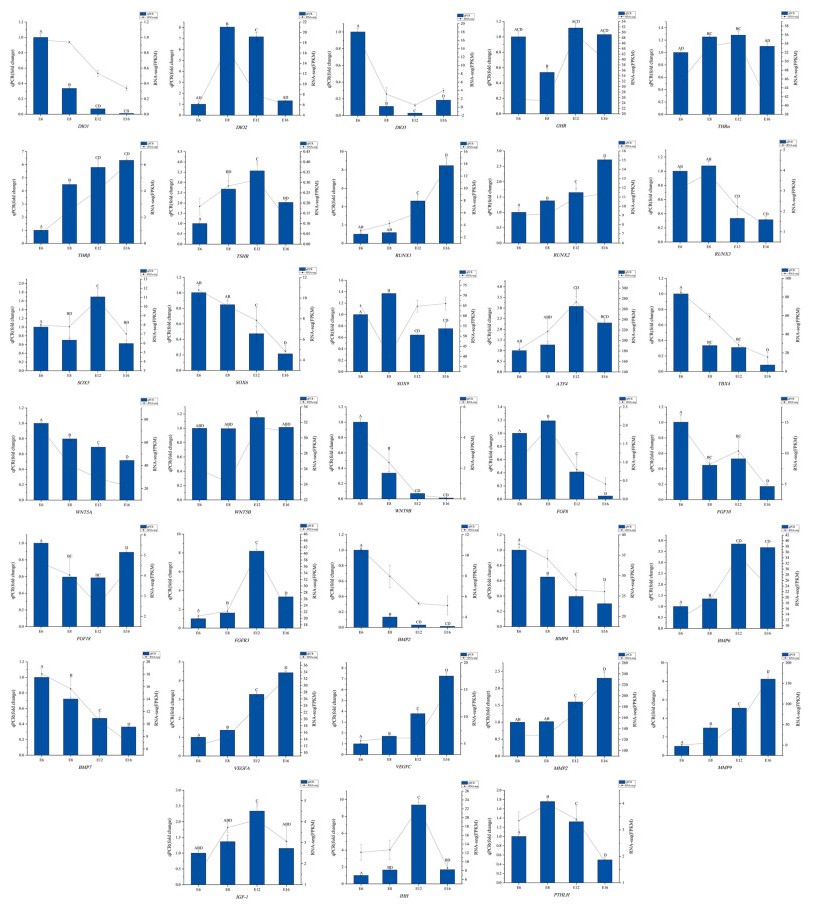
To verify the results of RNA-Seq data, 33 genes related to hindlimb bone formation were selected as candidate genes for qPCR assay. Table 2 summarizes the expression of 33 DEGs at 4 stages. Figure 6 showed that the expression patterns of 33 genes were generally consistent with RNA-Seq data. We found that various signaling factors were involved in the proliferation and differentiation of chondrocytes and osteoblasts with the development of the hindlimb bone of quail embryos.
| ID | Gene | Product | FPKM | |||
| E6 | E8 | E12 | E16 | |||
| #Systemic factors | ||||||
| 107317722 | DIO1 | type Ⅰ iodothyronine deiodinase isoform ×1 | 0.968 | 0.938 | 0.531 | 0.339 |
| 107315160 | DIO2 | type Ⅱ iodothyronine deiodinase isoform ×1 | 6.133 | 16.913 | 7.85 | 6.153 |
| 107315336 | DIO3 | thyroxine 5-deiodinase | 17.096 | 3.133 | 0.479 | 3.964 |
| 107305958 | GHR | growth hormone receptor | 25.265 | 24.914 | 49.257 | 40.078 |
| 107325239 | THRA | thyroid hormone receptor alpha | 45.368 | 53.452 | 54.457 | 41.054 |
| 107309297 | THRB | thyroid hormone receptor beta isoform ×1 | 0.75 | 2.531 | 4.075 | 5.974 |
| 107315169 | TSHR | thyrotropin receptor isoform ×1 | 0.184 | 0.283 | 0.314 | 0.127 |
| #Transcription factors | ||||||
| 107323234 | RUNX1 | runt-related transcription factor 1 isoform ×1 | 3.056 | 4.23 | 5.992 | 13.521 |
| 107312455 | RUNX2 | runt-related transcription factor 2 isoform ×1 | 9.031 | 9.114 | 10.944 | 11.366 |
| 107324053 | RUNX3 | runt-related transcription factor 3 | 3.116 | 3.879 | 2.229 | 1.297 |
| 107316728 | SOX5 | transcription factor SOX-5 isoform ×1 | 7.989 | 7.803 | 10.777 | 7.022 |
| 107314636 | SOX6 | transcription factor SOX-6 | 10.777 | 9.24 | 7.841 | 4.85 |
| 107322214 | SOX9 | transcription factor SOX-9 | 64.983 | 40.272 | 64.716 | 66.059 |
| 107312723 | ATF4 | transcription factor ATF4 | 183.225 | 217.37 | 273.667 | 224.673 |
| 107322512 | TBX4 | T-box transcription factor TBX4 | 85.041 | 58.916 | 28.497 | 15.48 |
| #Locally produced factors | ||||||
| 107320001 | WNT5A | protein Wnt-5a isoform ×1 | 72.549 | 40.459 | 28.899 | 22.444 |
| 107315234 | WNT5B | protein Wnt-5b | 25.89 | 24.287 | 31.373 | 30.867 |
| 107324983 | WNT9B | protein Wnt-9b | 4.025 | 2.378 | 0.242 | 0.072 |
| 107315948 | FGF8 | fibroblast growth factor 8 isoform ×1 | 1.063 | 1.951 | 0.8 | 0.409 |
| 107305970 | FGF10 | fibroblast growth factor 10 | 14.653 | 8.232 | 10.412 | 4.557 |
| 107320505 | FGF18 | fibroblast growth factor 18 isoform ×1 | 4.634 | 4.04 | 2.542 | 4.278 |
| 107313983 | FGFR3 | fibroblast growth factor receptor 3 | 20.727 | 22.248 | 39.97 | 23.058 |
| 107310979 | BMP2 | bone morphogenetic protein 2 | 10.592 | 7.968 | 5.323 | 5.131 |
| 107315532 | BMP4 | bone morphogenetic protein 4 | 37.709 | 34.134 | 26.489 | 26.085 |
| 107309636 | BMP6 | bone morphogenetic protein 6 | 12.682 | 18.87 | 34.771 | 23.962 |
| 107323076 | BMP7 | bone morphogenetic protein 7 | 18.08 | 15.655 | 10.492 | 7.018 |
| 109311307 | VEGFA | vascular endothelial growth factor A isoform ×1 | 12.133 | 14.675 | 23.638 | 32.107 |
| 107313491 | VEGFC | vascular endothelial growth factor C | 5.559 | 6.04 | 6.105 | 14.625 |
| 107319190 | MMP2 | matrix metalloproteinase-2 | 127.759 | 128.291 | 168.191 | 223.967 |
| 107323128 | MMP9 | matrix metalloproteinase-9 | 0.899 | 5.054 | 54.942 | 159.075 |
| 107316281 | IGF1 | Insulin-like growth factor Ⅰ | 1.514 | 3.725 | 4.093 | 3.063 |
| 107317031 | IHH | indian hedgehog protein | 12.138 | 12.714 | 22.267 | 8.602 |
| 107317927 | PTHLH | parathyroid hormone-related protein | 3.348 | 3.985 | 3.402 | 1.81 |
We selected seven systemic factors: type Ⅰ iodothyronine deiodinase (DIO1), type Ⅱ iodothyronine deiodinase (DIO2) and type Ⅲ iodothyronine deiodinase (DIO3), growth hormone receptor (GHR), thyroid hormone receptor alpha (THRα), thyroid hormone receptor beta (THRβ), thyroid stimulating hormone receptor (TSHR) and these genes were all involved in thyroid hormone signaling pathway. The mRNA expression of DIO1 and DIO3 was mainly expressed in the early stage of embryonic development and then was almost continuously significantly decreased from E6 to E16 while the mRNA expression of THRβ kept increasing from E6 to E16. The expression of DIO2, THRα and TSHR was increased at E8 and E12 then decreased at E16. GHR was decreased only at E8 and was highly expressed in the late stage of development.
The main transcription factors were RUNX family transcription factor 1 (RUNX1), RUNX family transcription factor 2 (RUNX2), RUNX family transcription factor 3 (RUNX3), SRY-box transcription factor 5 (SOX5), SRY-box transcription factor 6 (SOX6), SRY-box transcription factor 9 (SOX9), activating transcription factor 4 (ATF4) and T-box transcription factor 4 (TBX4). The transcriptional level of RUNX1, RUNX2 and ATF4 was kept increasing from E6 to E16 while that of RUNX3, SOX6 and TBX4 kept decreasing. The expression of SOX5 was significantly upregulated at E12, while the expression of SOX9 was upregulated at E8 and decreased at E12 and E16.
We selected 18 local secretory factors, including WNTs [WNT family member 5A (WNT5A), WNT family member 5B (WNT5B), WNT family member 9B (WNT9B)], fibroblast growth factors (FGFs) [FGF8, FGF10, FGF18, fibroblast growth factor receptor 3 (FGFR3)], bone morphogenetic proteins (BMPs) (BMP2, BMP4, BMP6, BMP7), vascular endothelial growth factors (VEGFs) (VEGFA and VEGFC), matrix metalloproteinases (MMPs) (MMP2 and MMP9), insulin-like growth factor 1 (IGF1), indian hedgehog (IHH) and parathyroid hormone like hormone (PTHLH). The mRNA level of WNT5A, FGF10, BMP2, BMP4 and BMP7 was almost continuously significantly decreased from E6 to E16 while VEGFA, VEGFC, BMP6, MMP2, MMP9 and IHH kept increasing from E6 to E16. FGFR3 and PTHLH was increased at E8 and E12 then decreased at E16. The expression of WNT5B and IGF1 was significantly upregulated at E12. The expression of FGF18 was decreased at E8 and E12 then increased at E16.
Japanese Quails especially domesticated varieties are primarily ground-dwelling birds, and the growth and development of hindlimb are important to provide the power needed for walking, running, and taking off during flight. Thus, the hindlimb is one of the earliest areas of ossification in the quail skeleton (Nakamura et al., 2019). Our study showed that cartilage ossification was observed at E6 in the femur and tibiofibula, then endochondral ossification of hindlimb gradually increased during embryonic development and most of hindlimb was ossified before hatching, which was consistent with previous studies (Pourlis et al., 1998; Pourlis and Antonopoulos, 2014). Moreover, transcriptomic results showed lots of signaling pathways were involved in regulating endochondral ossification of hindlimb, and the mRNA expression of genes changed with hindlimb bone development in quail embryo.
Thyroid hormone (TH) signaling pathway plays a key role in bone development and growth, and bone is a very sensitive and typical target tissue for T3 (Bassett and Williams, 2016). Thyroid stimulating hormone (TSH) is a hormone secreted by the pituitary gland to promote thyroid growth and function, which is conducive to the synthesis and release of thyroid hormones. Its receptor can receive thyroid stimulating hormone signal and accelerate chondrocyte hypertrophy. The expression of TSHR was highest at E12, also indicating that the high expression of TSH could promote the hypertrophy of chondrocytes. Moreover, TSH stimulates the thyroid gland to produce and release T3 and T4. T3 is a critical hormone for bone development, maintenance, and remodeling. It influences osteoblasts, osteoclasts, and bone matrix proteins, ensuring the proper formation, mineralization, and turnover of bone tissue (Darras et al., 2006). Because T3 and T4 have different concentrations and affinity in the tissue, T4 must be converted to T3 to function. T3 is mainly produced by T4 through the deiodination of three deiodinases (Bassett and Williams, 2016). DIO1 and DIO2 can convert T4 to T3 to increase TH activity, while DIO3 can catalyze T3 and T4 to generate inactive rT3 to decrease TH activity (Bo et al., 2018). In this study, the mRNA levels of three deiodinases were highly expressed in the early stage of bone development. For example, DIO1 and DIO3 had the highest expression at E6, and DIO2 had the highest expression at E8. The high expression of deiodinases might lead to increasing T3, which was conducive to the endochondral ossification of hindlimb. In addition, different expressions of thyroid hormone receptors (THRα and THRβ) also have a significant effect on endochondral ossification. Thyroid hormone receptors are present in bone cells and play a crucial role in regulating bone development, maintenance, and remodeling. Thyroid hormones, when bound to these receptors, influence the balance between bone formation and bone resorption, impact bone matrix composition, and help maintain bone density (Darras et al., 2006). In this study, the expression level of THRα was high in four developmental stages, and the expression level of THRβ increased gradually in the development process, which indicated that thyroid hormone receptors are important to endochondral ossification throughout the whole developmental process.
Moreover, growth hormone (GH) is very important for regulating bone homeostasis and can directly act on bone cells to promote longitudinal bone growth. GH also stimulates the expression of bone morphogenetic protein, which is very important for bone formation (Giustina et al., 2008). Hence, GHR expression was highest at E12, which was a critical period for long bones and was important for bone development. In addition, insulin-like growth factor 1 (IGF-1) is a peptide hormone that plays a significant role in bone development, growth, and maintenance. It is produced by the liver and other tissues in response to GH stimulation. IGF-1 is a key mediator of the effects of GH (Bo et al., 2018). Our study found that the expression of IGF1 was highest at E12, which was also consistent with the expression of GHR, and the high expression of IGF1 also indicated that it could enhance endochondral osteogenesis (Yakar et al., 2002).
In addition, several transcription factors were involved in endochondral ossification. The SOX (SRY-related HMG-box) family of transcription factors is one group of proteins that play essential roles in bone formation with SOX9 and SOX5/6 being particularly important. SOX9 is a critical transcription factor involved in chondrogenesis and it is essential for the differentiation of mesenchymal stem cells into chondrocytes and endochondral ossification (Gao et al., 2018; Lefebvre et al., 2019). In this study, the expression of SOX9 was firstly increased and then decreased, which was consistent with the rule of chondrocyte development and osteogenesis. Meanwhile, SOX9 is essential for chondrocyte differentiation and can work in synergy with SOX5 and SOX6 to ensure normal development of cartilage primordia (Lefebvre, 2019). Previous studies showed that SOX5 and SOX6 knockout mice suffered from severe systemic chondroplasia (Smits et al., 2001). Our study found that SOX5 and SOX6 expression gradually decreased during development, which may indicate that they were more involved in chondrocyte proliferation than hypertrophy (Provot and Schipani, 2005).
Moreover, the RUNX family of transcription factors consists of three members: RUNX1, RUNX2, and RUNX3, and they all play a pivotal role in bone development. RUNX1 mediates chondrogenesis and chondrocyte differentiation, while RUNX2 promotes end-stage chondrocyte maturation and mineralization. RUNX1 is highly expressed in chondrocytes and osteoblasts (Kimura et al., 2010). RUNX1 and RUNX2 are involved in key steps in the differentiation of mesenchymal stem cells into chondrocytes (Wang et al., 2005). Our results showed that the expression level of RUNX1 increased gradually during quail embryonic hindlimb development, which may indicate that RUNX1 played a role not only in inducing mesenchymal stem cells in early bone development, but also in vascular formation in late bone development. Besides, RUNX3 is expressed in prehypertrophic chondrocytes and may induce early chondrogenesis through synergistic effect with RUNX1 (Soung et al., 2007). RUNX2 co-induces chondrocyte hypertrophy with RUNX3, and the absence of RUNX2 caused delayed chondrocyte hypertrophy in mice (Provot and Schipani, 2005; Gao et al., 2018). Our results suggested that continued upregulation of RUNX2 was beneficial for maintaining chondrocyte proliferation and hypertrophy as well as chondrocyte vascular invasion. The expression regulation trend of RUNX3 was opposite to that of RUNX2, indicating that RUNX3 mainly played a role in regulating chondrocyte hypertrophy at the early stage of bone development.
Meanwhile, RUNX2 regulates the expression of activating transcription factor 4 (ATF4) and ATF4 in turn amplifies the expression of genes necessary for later stages of osteoblast differentiation and mineralization. ATF4 is a member of the ATF/CREB family of basic leucine zipper (bZIP) transcription factors and it is known to be a key regulator of bone formation. In particular, ATF4 is a critical transcription factor and it regulates osteoblast differentiation, bone matrix production and mineralization (Zhu et al., 2013). Both ATF4 and RUNX2 are transcription factors that are crucial for osteoblast differentiation and bone development. RUNX2 is involved in the early commitment of mesenchymal stem cells to the osteoblast lineage, while ATF4 plays a role in later stages of osteoblast differentiation and bone matrix mineralization (Yu et al., 2013). The expression of ATF4 was highest at E12 suggesting its role in late bone development of quail embryo.
Meanwhile, T-box transcription factors (TBX) are a family of transcription factors that play crucial roles in the regulation of gene expression and are involved in various developmental processes including bone development. TBX4 is mainly involved in the hindlimb development of chicken embryos, and also controls the initiation of limb bud development and the expression of limb bud mesenchymal in the hindlimb of quail embryos (Tickle, 2015). In this study, the expression level of TBX4 decreased gradually with the development of hindlimb, and reached the highest level at the early stage of embryonic development, emphasizing that TBX4 played an essential role in initiating hindlimb buds and promoting early bone development.
A lot of secretory factors were also associated with bone development. Bone morphogenetic proteins (BMPs) are a group of growth factors that belong to transforming growth factor-beta (TGF-β) superfamily. BMPs are critical regulators of bone development, influencing the differentiation of progenitor cells into bone-forming osteoblasts and promoting bone formation and repair. They play pivotal roles in various developmental processes, including bone and cartilage formation. BMPs were originally identified by their ability to induce bone formation (Long and Ornitz, 2013; Gao et al., 2018). BMP2, BMP4 and BMP7 are all potent inducers of osteogenesis during embryonic development. They help initiate the formation of bones and joints and stimulate the differentiation of mesenchymal cells into chondrocytes during early development (Bandyopadhyay et al., 2006). Our study also showed that the expression levels of BMP2, BMP4, and BMP7 decreased with hindlimb development, indicating that these genes mainly played an important role in the stage of chondrocyte proliferation. While the expression level of BMP6 gradually increased and reached the maximum at E12, and it helped ensure the proper development and growth of bones by orchestrating the transformation of cartilage into bone during endochondral ossification.
Besides, FGFs and their receptors are critical regulators of bone development and homeostasis. FGF/FGFR signaling can influence various processes, from chondrocyte differentiation in the growth plate to osteoblast and osteoclast activity in mature bone (Marie et al., 2012). FGF8 played an important role in limb development and also regulates chondrogenic differentiation (Su et al., 2014). FGF10 acted upstream of FGF8 and was a limb bud initiator expressed in mesenchymal tissue. For instance, mice of knocked-out FGF10 completely lost their forelimbs and hindlimbs (Su et al., 2014). The FGF8 mutation also seriously affected limb development in mice, and some bone elements could not be produced normally (Boulet et al., 2004). Our study showed that the expression of FGF10 was highest in E6, which was also the period when ossification centers appeared in the hindlimbs, and a large number of mesenchymal cells were needed to gather to begin limb development. FGF10 acted on the upstream of FGF8, and we also observed the highest expression of FGF8 at E8, suggesting that FGF10 initiated a large number of mesenchymal cells to participate in bone formation, and then transmitted signals to FGF8 to continue to participate in limb development. Moreover, FGF18 is a key regulator involved in chondrogenesis and angiogenesis during early bone development. FGF18 negatively regulated chondrocyte proliferation and differentiation through FGFR3, and our study also showed that the expression trend of FGF18 was opposite to that of FGFR3. FGFR3 promoted chondrocyte proliferation during early embryonic development, and in turn inhibited chondrocyte proliferation and hypertrophic differentiation during late embryonic development (Liu et al., 2007).
In addition, the WNT signaling pathway plays an important role in regulating bone development and maintaining homeostasis (Baron and Rawadi, 2007). A large number of WNT signaling molecules are expressed in growing cartilage (Zhong et al., 2014; Bo et al., 2018). Among them, WNT5A has been widely studied and plays a key role in embryonic bone development by activating osteoclast precursor surface receptors to enhance osteoclast production (Zhong et al., 2014). In addition, WNT5A can promote osteoblast differentiation and enhance bone formation (Lerner and Ohlsson, 2015; Kobayashi et al., 2016). Meanwhile, WNT5B is expressed in mesenchymal stem cells, chondrocytes, osteoblasts and osteoclasts. In long bones, it is mainly expressed in the pre-hypertrophic region and activates chondrocyte proliferation (Suthon et al., 2021). WNT5A and WNT5B signaling are necessary for chondrocyte hypertrophy (Long and Ornitz, 2013). Our study showed that the expression of WNT5A decreased, indicating that WNT5A plays an important role in the stage of chondrocyte proliferation and hypertrophy. Our data showed that WNT5B expression did not change significantly across four stages, suggesting that WNT5B played an important role in each key stage of bone formation to promote bone formation in different types of osteocytes. Moreover, the expression of WNT9B also decreased during the developmental process and it helped stimulate the differentiation of mesenchymal cells into chondrocytes and these chondrocytes formed the cartilaginous template that served as a precursor to bone during embryonic development.
Furthermore, matrix metalloproteinase (MMP) family is involved in the process of matrix invasion and bone formation during the normal development of bone, and it is a key factor in the initiation of angiogenesis (Bo et al., 2018; Zhang et al., 2020). MMP2 and MMP9 can jointly stimulate angiogenesis (van Hinsbergh et al., 2006). Our study showed that the expression levels of MMP2 and MMP9 were gradually increased, indicating that these genes mainly promoted vascular development and chondrocyte invasion into the cellular matrix to form bone during late development.
Moreover, vascular endothelial growth factor (VEGF) is a critical signaling protein that plays a significant role in regulating angiogenesis and is closely associated with bone development and maintenance. Its ability to stimulate the growth of blood vessels is essential for providing the necessary nutrients and oxygen to bone tissue during development, growth, and repair (Hu and Olsen, 2016; Casie Chetty et al., 2017). Lack of VEGF caused delayed invasion of ossifying central vessels (Zelzer et al., 2002). VEGFA is an important growth factor regulating vascular development, and can also regulate terminally differentiated chondrocytes to participate in endochondral osteogenesis (Hu and Olsen, 2016). Our study showed that the expression level of VEGFA gradually increased with hindlimb development, implying that vascular invasion aggravated at late developmental stage and promoted the transformation of osteoblasts into bone. Moreover, VEGFC was expressed in almost all tissues at all stages of embryonic development, and overexpression in bone can promote the formation of lymphocytes in bone and osteoclast-mediated bone resorption (Hominick et al., 2018). The expression of VEGFC reached a peak in late development, similar to VEGFA, which promoted the development of blood vessels in cartilage and bone. Therefore, we concluded that MMP2, MMP9, VEGFA and VEGFC could stimulate angiogenesis and cartilage matrix invasion.
Meanwhile, Indian hedgehog (IHH) and parathyroid hormone-related protein (PTHrP), which is encoded by the PTHLH gene, are two key signaling molecules that play crucial roles in the process of endochondral ossification (Long et al., 2004; Gao et al., 2018). Indian hedgehog, a member of the hedgehog family of proteins, is produced by pre-hypertrophic and hypertrophic chondrocytes within the cartilage growth plate. It acts as a signaling molecule that promotes the proliferation of chondrocytes, especially in the proliferative and pre-hypertrophic zones of the growth plate. Our results showed that the expression of IHH first increased and then decreased, and reached the highest expression at E12, which also indicated that IHH played an important role in promoting chondrocyte proliferation and differentiation. Moreover, PTHrP is produced by mature chondrocytes located in the upper zone of growth plate. PTHrP helps maintain chondrocyte proliferation by preventing their premature differentiation into hypertrophic chondrocytes. Accordingly, the mRNA expression of PTHLH reached its maximum at E8, which may inhibit the differentiation of chondrocytes into hypertrophic chondrocytes and ensure proper bone growth and elongation.
The present study showed that the hindlimb developed from cartilage into hard bone and became increasingly ossified during the process of embryonic development. RNA-Seq analysis showed that various signaling pathways were involved in the proliferation and differentiation of chondrocytes and osteoblasts with the development of hindlimb bone of quail embryos. The study may benefit understanding of the molecular regulation of bone development and endochondral ossification.
Xuan Li: Conceptualization, Data curation, Investigation, Methodology, Validation, Writing – original draft. Yuxin Zhang: Formal analysis, Methodology, Visualization, Writing – review & editing. Hongfeng Zhao: Conceptualization, Data curation, Methodology, Supervision, Writing – review & editing, Funding acquisition, Writing – original draft.
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
We are grateful to Ms. Lingyang Jing and Qingyu Zhang for valuable comments.
Supplementary data to this article can be found online at https://doi.org/10.1016/j.avrs.2023.100152.
Genes: ATF4: activating transcription factor 4
BMP2: bone morphogenetic protein 2
BMP4: bone morphogenetic protein 4
BMP6: bone morphogenetic protein 6
BMP7: bone morphogenetic protein 7
DIO1: type Ⅰ iodothyronine deiodinase
DIO2: type Ⅱ iodothyronine deiodinase
DIO3: type Ⅲ iodothyronine deiodinase
FGF8: fibroblast growth factor 8
FGF10: fibroblast growth factor 10
FGF18: fibroblast growth factor 18
FGFR3: fibroblast growth factor receptor 3
GHR: growth hormone receptor
IGF-1: insulin like growth factor 1
IHH: Indian hedgehog signaling molecule
MMP2: matrix metallopeptidase 2
MMP9: matrix metallopeptidase 9
PTHrP: parathyroid hormone-related protein
PTHLH: parathyroid hormone like hormone
RUNX1: RUNX family transcription factor 1
RUNX2: RUNX family transcription factor 2
RUNX3: RUNX family transcription factor 3
SOX5: SRY-box transcription factor 5
SOX6: SRY-box transcription factor 5
SOX9: SRY-box transcription factor 9
TBX4: T-box transcription factor 4
THRα: thyroid hormone receptor alpha
THRβ: thyroid hormone receptor beta
TSHR: thyroid stimulating hormone receptor
VEGFA: vascular endothelial growth factor A
VEGFC: vascular endothelial growth factor C
WNT5A: WNT family member 5A
WNT5B: WNT family member 5B
WNT9B: WNT family member 9B
|
Baer, J., Lansford, R., Cheng, K., 2015. Chapter 22 – Japanese quail as a laboratory animal model. In: Fox, J.G., Anderson, L.C., Otto, G.M., Pritchett-Corning, K.R., Whary, M.T. (Eds.), Laboratory Animal Medicine, third ed. American College of Laboratory Animal Medicine, Academic Press, pp. 1087–1108.
|
|
Livak, K.J., Schmittgen, T.D., 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402-408.
|
|
Maxwell, E. E. (2008). Comparative embryonic development of the skeleton of the domestic turkey (Meleagris gallopavo) and other galliform birds. Zoology, 111(3), 242-257.
|
|
Morris, K.M., Hindle, M.M., Boitard, S., Burt, D.W., Danner, A.F., Eory, L., et al., 2020. The quail genome: insights into social behaviour, seasonal biology and infectious disease response. BMC Biol. 18, 14.
|
|
Wang, Z., Gerstein, M., Snyder, M., 2009. RNA-Seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57-63.
|
|
Wang, L., Jing, L., Zhang, Q., Li, S., Wang, Y., Zhao, H., 2022. Lead induced thymic immunosuppression in Japanese quail (Coturnix japonica) via oxidative stress-based T cell receptor pathway signaling inhibition. J. Inorg. Biochem. 235, 111950.
|
|
Zhang, G., Zhang, Y., Jing, L., Zhao, H., 2023. Lead exposure induced developmental nephrotoxicity in Japanese quail (Coturnix japonica) via oxidative stress-based PI3K/AKT pathway inhibition and NF-κB pathway activation. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 268, 109599.
|
| Developmental stage | Femur length (mm, mean ± SE) | Hindlimb length (mm, mean ± SE) | Ossified proportion of femur (%) |
| E6 | 1.64 ± 0.28 | 9.31 ± 0.06 | 20.6 ± 2.2 |
| E7 | 3.15 ± 0.25 | 11.57 ± 0.13 | 28.6 ± 2.6 |
| E8 | 3.82 ± 0.14 | 12.81 ± 0.13 | 32.8 ± 2.4 |
| E9 | 5.27 ± 0.27 | 20.14 ± 0.34 | 52.6 ± 3.6 |
| E10 | 5.99 ± 0.12 | 23.44 ± 0.18 | 61.4 ± 1.6 |
| E11 | 6.39 ± 0.38 | 29.36 ± 0.14 | 64.2 ± 2.2 |
| E12 | 6.98 ± 0.04 | 33.64 ± 0.27 | 70.8 ± 2.4 |
| E13 | 9.33 ± 0.31 | 40.04 ± 0.06 | 75.2 ± 3.2 |
| E14 | 10.57 ± 0.06 | 44.17 ± 0.14 | 79.4 ± 2.2 |
| E15 | 11.78 ± 0.12 | 51.16 ± 0.39 | 81.6 ± 3.4 |
| E16 | 13.27 ± 0.05 | 54.81 ± 0.38 | 86.2 ± 1.8 |
| ID | Gene | Product | FPKM | |||
| E6 | E8 | E12 | E16 | |||
| #Systemic factors | ||||||
| 107317722 | DIO1 | type Ⅰ iodothyronine deiodinase isoform ×1 | 0.968 | 0.938 | 0.531 | 0.339 |
| 107315160 | DIO2 | type Ⅱ iodothyronine deiodinase isoform ×1 | 6.133 | 16.913 | 7.85 | 6.153 |
| 107315336 | DIO3 | thyroxine 5-deiodinase | 17.096 | 3.133 | 0.479 | 3.964 |
| 107305958 | GHR | growth hormone receptor | 25.265 | 24.914 | 49.257 | 40.078 |
| 107325239 | THRA | thyroid hormone receptor alpha | 45.368 | 53.452 | 54.457 | 41.054 |
| 107309297 | THRB | thyroid hormone receptor beta isoform ×1 | 0.75 | 2.531 | 4.075 | 5.974 |
| 107315169 | TSHR | thyrotropin receptor isoform ×1 | 0.184 | 0.283 | 0.314 | 0.127 |
| #Transcription factors | ||||||
| 107323234 | RUNX1 | runt-related transcription factor 1 isoform ×1 | 3.056 | 4.23 | 5.992 | 13.521 |
| 107312455 | RUNX2 | runt-related transcription factor 2 isoform ×1 | 9.031 | 9.114 | 10.944 | 11.366 |
| 107324053 | RUNX3 | runt-related transcription factor 3 | 3.116 | 3.879 | 2.229 | 1.297 |
| 107316728 | SOX5 | transcription factor SOX-5 isoform ×1 | 7.989 | 7.803 | 10.777 | 7.022 |
| 107314636 | SOX6 | transcription factor SOX-6 | 10.777 | 9.24 | 7.841 | 4.85 |
| 107322214 | SOX9 | transcription factor SOX-9 | 64.983 | 40.272 | 64.716 | 66.059 |
| 107312723 | ATF4 | transcription factor ATF4 | 183.225 | 217.37 | 273.667 | 224.673 |
| 107322512 | TBX4 | T-box transcription factor TBX4 | 85.041 | 58.916 | 28.497 | 15.48 |
| #Locally produced factors | ||||||
| 107320001 | WNT5A | protein Wnt-5a isoform ×1 | 72.549 | 40.459 | 28.899 | 22.444 |
| 107315234 | WNT5B | protein Wnt-5b | 25.89 | 24.287 | 31.373 | 30.867 |
| 107324983 | WNT9B | protein Wnt-9b | 4.025 | 2.378 | 0.242 | 0.072 |
| 107315948 | FGF8 | fibroblast growth factor 8 isoform ×1 | 1.063 | 1.951 | 0.8 | 0.409 |
| 107305970 | FGF10 | fibroblast growth factor 10 | 14.653 | 8.232 | 10.412 | 4.557 |
| 107320505 | FGF18 | fibroblast growth factor 18 isoform ×1 | 4.634 | 4.04 | 2.542 | 4.278 |
| 107313983 | FGFR3 | fibroblast growth factor receptor 3 | 20.727 | 22.248 | 39.97 | 23.058 |
| 107310979 | BMP2 | bone morphogenetic protein 2 | 10.592 | 7.968 | 5.323 | 5.131 |
| 107315532 | BMP4 | bone morphogenetic protein 4 | 37.709 | 34.134 | 26.489 | 26.085 |
| 107309636 | BMP6 | bone morphogenetic protein 6 | 12.682 | 18.87 | 34.771 | 23.962 |
| 107323076 | BMP7 | bone morphogenetic protein 7 | 18.08 | 15.655 | 10.492 | 7.018 |
| 109311307 | VEGFA | vascular endothelial growth factor A isoform ×1 | 12.133 | 14.675 | 23.638 | 32.107 |
| 107313491 | VEGFC | vascular endothelial growth factor C | 5.559 | 6.04 | 6.105 | 14.625 |
| 107319190 | MMP2 | matrix metalloproteinase-2 | 127.759 | 128.291 | 168.191 | 223.967 |
| 107323128 | MMP9 | matrix metalloproteinase-9 | 0.899 | 5.054 | 54.942 | 159.075 |
| 107316281 | IGF1 | Insulin-like growth factor Ⅰ | 1.514 | 3.725 | 4.093 | 3.063 |
| 107317031 | IHH | indian hedgehog protein | 12.138 | 12.714 | 22.267 | 8.602 |
| 107317927 | PTHLH | parathyroid hormone-related protein | 3.348 | 3.985 | 3.402 | 1.81 |